Three dimensional structures
(38 PDB entries- in collaboration with the Tanner group at the University of Missouri)
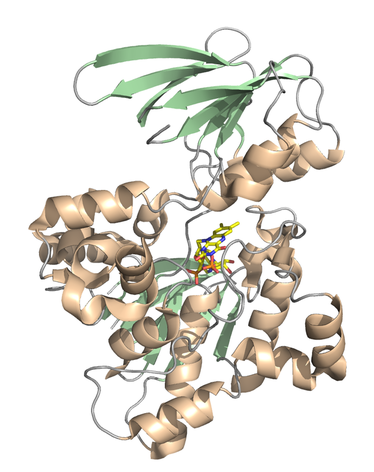
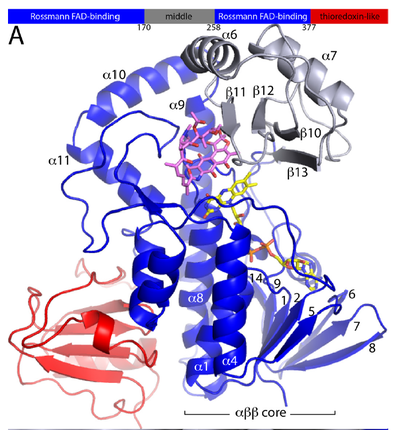
Structure of allyl-mercarptan monooxygenase from Allium satuvum (garlic)
6WPU- FAD bound
|
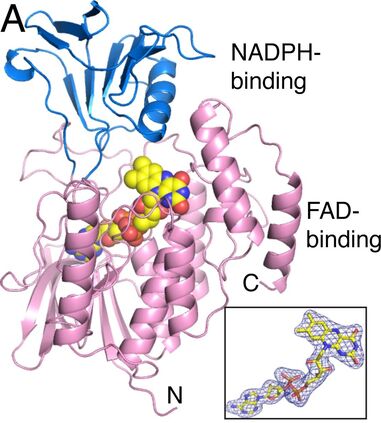
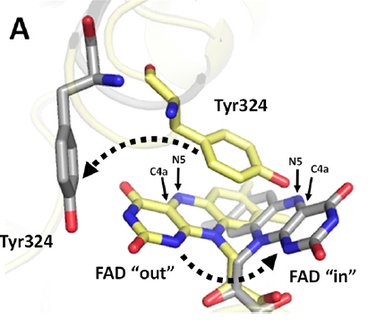
Structures of N-monooxygenases (NMO)
A) Structures of N5-ornithine monooxygenase from Aspergillus fumigatus
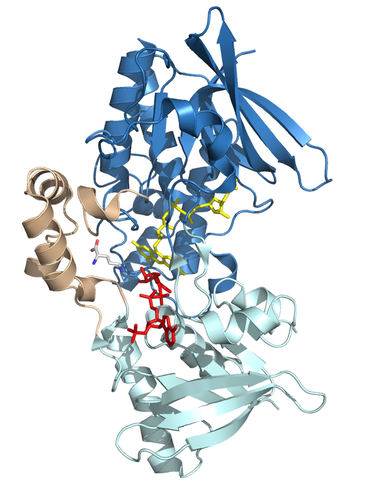
4B63- in complex with NADP and ornithine
4B64- in complex with NADP and lysine
4B65- in complex with NADP (in the reduced form)
4b66- in complex with Arg and NADP (in the reduced form)
4B67- in complex with NADP and ornithine (re-oxidized)
4B68- in complex with NADP and Arg (re-oxidized)
4B69- in complex with ornithine
4NZH- R279A mutant
5CKU- N323A mutant
6X0H Ligand free oxidized
6X0I oxidized with NADP+
6X0J reduced with NADP+ and ornithine
6X0K reduced with ornithine
4B64- in complex with NADP and lysine
4B65- in complex with NADP (in the reduced form)
4b66- in complex with Arg and NADP (in the reduced form)
4B67- in complex with NADP and ornithine (re-oxidized)
4B68- in complex with NADP and Arg (re-oxidized)
4B69- in complex with ornithine
4NZH- R279A mutant
5CKU- N323A mutant
6X0H Ligand free oxidized
6X0I oxidized with NADP+
6X0J reduced with NADP+ and ornithine
6X0K reduced with ornithine
B) Structure of N6-Lysine monooxygenase from Nocardia farcinica
C) Structure of putrescine monooxygenase from Acinetobacter baumannii
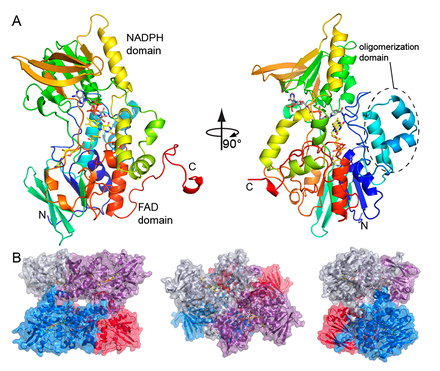
Structures of Rifampicin monooxygenase (RifMO)
Structures of UDP-galactopyranose mutases (UGMs)
A) UGM from A. fumigatus
|
5HHF- FAD-Galactopyranose intermediate
3UTE- in complex with sulfate 3UTF- in the reduced state 3UTG- in complex with UDP 3UTH- in complex with UDP-Galp (in the reduced state) 4GDC- in complex with NADPH 4GDD- in complex with NADH 4GDE- NADPH- reduced 4U8I- F66A ligand free 4U8J- Y104A ligand free 4U8K- Q107A ligand free 4U8L- N207A ligand free 4U8M- Y317A ligand free 4U8N-F66A in complex with UDP 4WX1-F66A in complex with UDP-Galp 4U8O-N207A in complex with UDP 4U8P-Y317A- in complex with UDP |